r-lequin
This is an old revision of the document!
Table of Contents
R-lequin
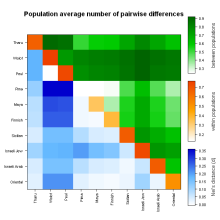
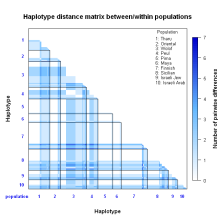
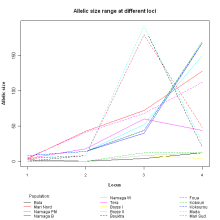
R-lequin is an R interface for the graphical display of Arlequin outputs. It should add value and help interpreting results from large Arlequin analyses.
R code for graphics
Documentations:
lines:
- Haplotype:
- expected/observed haplotype (with CI)
- save graphic as a .png file
- Allelic size range (lines with different colors)
matrix:
-
- read data from a HTML file
- read data from a XML file
- move legend
- haplotype distance:
- haplotype distance (line by line input)
- improved: matrix rotated, more labels
-
- with mixed data from several populations
- multiple plots in one graphic:
bar:
points:
XML:
-
- half matrix data
- full matrix data
- table data
Arlequin output file changes (HTML to XML)
Delete:
- <html>, <head>, <style>, <body> and <PRE> tags
- the space ( ) in the empty <A> tag or replace it with the space code: 
O;
Insert:
- XML declaration on top of the output file:
<?xml version="1.0" encoding="ISO-8859-1"?> <?xml-stylesheet type="text/xsl" href="..."?>
- root tag
- tags around titles and other text which should be specially formatted
- tags around data (respectively around data which should be displayed with no special format)
- specific tags around data which is needed to make graphics in R:
- unique names for a special kind of data (the format of the data within this tag must always be the same)
- no sample names/population names with wihte space, use '_' instead
example:
tutorials
links
r-lequin.1228496003.txt.gz · Last modified: 2008/12/05 17:53 by heidi
